- mRNA Sequencing
- Small RNA Sequencing
- Whole Transcriptome Sequencing
- Single Cell RNA Sequencing
- circularRNA Sequencing
- DrSeq GEScore Biostimulants Testing
mRNA Sequencing
Service Description: mRNA Sequencing on Illumina NovaSeq 6000 Platform
Welcome to Nucleome Informatics’ mRNA Sequencing service, powered by the cutting-edge Illumina NovaSeq 6000 platform. Our comprehensive mRNA sequencing service empowers researchers to unravel the complex transcriptome landscape and gain valuable insights into gene expression patterns. Whether you’re exploring gene regulation, biomarker discovery, or understanding disease mechanisms, our state-of-the-art technology and expertise will accelerate your research endeavors.
Option 1: Basic mRNA Sequencing Service
For researchers seeking essential mRNA sequencing and analysis
Sample Requirement:
- Sample Quantity: A minimum of 100 ng of high-quality RNA is required for successful library preparation.
- Sample Purity: RNA samples must have an RNA Integrity Number (RIN) of 7 or higher, ensuring minimal degradation and contamination.
- Sample Concentration: RNA samples should have a concentration of at least 10 ng/μl.
Workflow:
- RNA Isolation and Quality Control:
- Optional RNA isolation service is available if you require extraction and quality assessment of RNA samples.
- Alternatively, you can provide pre-isolated and quality-checked RNA samples.
- Library Preparation:
- State-of-the-art library preparation methods for constructing stranded RNA-seq libraries.
- cDNA synthesis, adapter ligation, and mRNA fragment enrichment steps.
- Library Quality Check:
- Stringent quality checks to assess size distribution, concentration, and adapter ligation efficiency.
- Sequencing:
- High-throughput, paired-end (2×150 bp) sequencing on the Illumina NovaSeq 6000 platform.
- Standard sequencing depth for most applications, with the option to customize according to project requirements.
- Data Analysis:
- Read Alignment to the reference genome or transcriptome.
- Differential Gene Expression Analysis to identify significant changes in gene expression.
- Functional Annotation and pathway analysis for functional insights.
- Comprehensive visualizations including heatmaps, volcano plots, and gene expression profiles.
Data Delivery:
- Comprehensive report with all relevant data, analysis results, and visualizations.
- Access to raw sequencing data and processed files through our secure online portal.
Option 2: Enhanced mRNA Sequencing Service
For researchers requiring more in-depth analysis and customized solutions
Additional Unique Points:
- In-depth consultation with our expert bioinformaticians to tailor the analysis based on specific research objectives.
- Ability to incorporate additional replicates or experimental conditions for robust statistical analysis.
- Advanced bioinformatics pipelines to uncover alternative splicing events and isoform quantification.
Option 3: Premium mRNA Sequencing Service
For large-scale projects and comprehensive transcriptome profiling
Additional Unique Points:
- Expedited turnaround time for urgent projects.
- Highly experienced project management team to handle large-scale studies effectively.
- Long-read sequencing options for challenging transcriptome analysis.
At Nucleome Informatics, we are committed to delivering high-quality results and exceptional customer service. Our experienced team of scientists and bioinformaticians is ready to assist you at every step of your mRNA sequencing project.
Unlock the mysteries of gene expression with our mRNA sequencing service on Illumina NovaSeq 6000. Contact us today to discuss your project requirements and to receive a customized quote.


Small RNA Sequencing
Small RNA Sequencing and miRNA Sequencing at Nucleome Informatics
Welcome to Nucleome Informatics, your leading partner in Small RNA Sequencing and miRNA Sequencing services. With over 10 years of extensive experience in this field, we offer cutting-edge technologies, a proven track record, and a diverse clientele that encompasses academic researchers, pharmaceutical companies, and biotechnology firms. Whether you’re exploring microRNA (miRNA) expression profiles, small RNA regulation, or biomarker discovery, our expertise will unlock the potential of your research.
Sample Requirement:
Our Small RNA Sequencing and miRNA Sequencing services accommodate various sample types, including total RNA and enriched small RNA samples. The following are the sample requirements for successful sequencing:
– Sample Quantity: A minimum of 500 ng of total RNA is required for Small RNA Sequencing and miRNA Sequencing library preparation.
– Sample Purity: RNA samples must have an RNA Integrity Number (RIN) of 7 or higher to ensure minimal degradation and contamination.
– Sample Concentration: RNA samples should have a concentration of at least 25 ng/μl.
Sequencing Details:
For Small RNA Sequencing and miRNA Sequencing, we utilize state-of-the-art next-generation sequencing platforms, providing the following read lengths and number of reads per sample:
– Read Length: Our standard sequencing offers read lengths of up to 150 base pairs (bp), enabling precise identification of small RNA species and detailed profiling of miRNA expression (SE50bp).
– Number of Reads per Sample: We ensure deep coverage of your small RNA samples with a minimum of 10 million reads per sample or more. This high read depth enhances the sensitivity and accuracy of miRNA expression analysis.
Workflow:
Our streamlined workflow ensures accurate and reliable results for Small RNA Sequencing and miRNA Sequencing:
1. RNA Isolation and Quality Control:
– Optional RNA isolation service is available to ensure high-quality RNA extraction.
– Rigorous quality control measures are employed to assess the integrity and purity of the RNA samples.
2. Small RNA Library Preparation:
– Specialized library preparation methods for constructing small RNA-seq libraries, including miRNAs and other small non-coding RNAs.
– Adapter ligation, reverse transcription, and size selection steps to capture a comprehensive range of small RNA species.
3. Library Quality Check:
– Stringent quality checks to evaluate library size distribution, concentration, and adapter ligation efficiency.
4. Sequencing:
– High-throughput sequencing on state-of-the-art platforms to ensure deep coverage and accurate profiling of small RNAs.
5. Data Analysis:
– Alignment to small RNA reference databases for miRNA identification and quantification.
– Differential Expression Analysis to uncover significant changes in miRNA expression between experimental conditions or sample groups.
– Pathway analysis to gain functional insights into miRNA-target interactions.
– Customized bioinformatics analysis based on specific research objectives.
Applications in Plant, Animal, Human, and Disease Research:
– Plant Research: Small RNA Sequencing and miRNA Sequencing are widely used in plant research to study gene regulation, stress responses, and the role of miRNAs in developmental processes.
– Animal Research: In animal research, these sequencing techniques are applied to investigate miRNA-mediated gene expression control, neurodevelopment, and disease mechanisms.
– Human Research: Small RNA Sequencing and miRNA Sequencing play a crucial role in human research, aiding in the discovery of disease biomarkers, studying miRNA involvement in cancer, and understanding miRNA-based therapeutics.
– Disease Research: Our services are instrumental in disease research to identify disease-specific miRNA signatures, unravel disease mechanisms, and discover potential therapeutic targets.
Discover the hidden potential of small RNA molecules with our Small RNA Sequencing and miRNA Sequencing services. Partner with Nucleome Informatics to propel your research to new heights.
*Contact us today to discuss your project requirements and benefit from our expertise in Small RNA Sequencing and miRNA Sequencing.*
Long Non-Coding RNA Sequencing
Long Non-Coding RNA (lncRNA) Sequencing with Ribo-zero Treatment Service at Nucleome Informatics
Welcome to Nucleome Informatics’ Long Non-Coding RNA (lncRNA) Sequencing service, where we provide cutting-edge technologies and expert analysis to unlock the vast potential of non-coding RNAs. With over 10 years of extensive experience in genomics research, our state-of-the-art service encompasses advanced techniques, including Ribo-zero treatment, which enhances the accuracy and sensitivity of lncRNA sequencing.
Sample Requirement: To ensure successful lncRNA sequencing with Ribo-zero treatment, we require high-quality total RNA or enriched lncRNA samples. The following are the sample requirements for optimal library preparation:
- Sample Quantity: A minimum of 200 ng of total RNA is needed for lncRNA sequencing library preparation.
- Sample Purity: RNA samples must have an RNA Integrity Number (RIN) of 7 or higher to ensure minimal degradation and contamination.
- Sample Concentration: RNA samples should have a concentration of at least 20 ng/μl.
Ribo-zero Treatment: Our lncRNA Sequencing service includes the application of Ribo-zero treatment during the library preparation step. Ribo-zero is a ribosomal RNA depletion technology that effectively removes rRNA molecules from the total RNA sample. By eliminating rRNA, we enrich the remaining RNA pool, including lncRNAs, small non-coding RNAs, and mRNA, which are crucial for studying gene expression regulation.
Library Preparation: Our expert team employs specialized library preparation methods to capture and enrich lncRNAs for accurate and comprehensive sequencing:
- RNA Isolation and Quality Control:
- We offer optional RNA isolation services to ensure high-quality RNA extraction.
- Rigorous quality control measures are employed to assess RNA integrity and purity.
- Ribo-zero Treatment:
- The Ribo-zero treatment is applied to selectively remove ribosomal RNA molecules from the total RNA sample.
- lncRNA Enrichment:
- After Ribo-zero treatment, we perform lncRNA enrichment to selectively capture non-coding RNA molecules, including long non-coding RNAs, for focused sequencing.
- Library Construction:
- Strand-specific library preparation to accurately determine the orientation of lncRNAs.
- cDNA synthesis, adapter ligation, and amplification steps are optimized for robust library construction.
- Library Quality Check:
- Stringent quality checks to evaluate library size distribution, concentration, and adapter ligation efficiency.
Sequencing on Illumina NovaSeq 6000: Our lncRNA Sequencing service utilizes the highly advanced Illumina NovaSeq 6000 platform, known for its superior sequencing capacity and cost-effectiveness. The sequencing process includes:
- Read Type: We offer paired-end (2×150 bp) sequencing for optimal coverage and accurate identification of lncRNAs.
- Sequencing Depth: Our standard sequencing depth ensures comprehensive coverage, with the flexibility to customize based on your specific research needs.
Data Analysis: Our bioinformatics experts meticulously analyze the lncRNA sequencing data, providing a comprehensive report with actionable insights:
- lncRNA Identification: Precise identification and annotation of lncRNAs from the sequencing data.
- Differential Expression Analysis: Comparative analysis to identify differentially expressed lncRNAs across experimental conditions or sample groups.
- Functional Annotation: Gene ontology and pathway analysis to uncover the biological functions of differentially expressed lncRNAs.
- Interaction Network Analysis: Exploration of lncRNA-mRNA interactions to understand regulatory networks.
Importance of lncRNA Sequencing in Research: Long Non-Coding RNAs play crucial roles in diverse biological processes, and their dysregulation has been implicated in numerous diseases. Our lncRNA Sequencing service empowers researchers in:
- Plant Research: Identifying lncRNAs involved in developmental processes, stress responses, and crop improvement.
- Animal Research: Exploring lncRNA-mediated gene regulation, embryonic development, and disease pathways.
- Human Research: Unraveling lncRNA signatures as potential biomarkers and therapeutic targets in various diseases, including cancer and neurodegenerative disorders.
- Disease Research: Understanding the roles of lncRNAs in disease pathogenesis, paving the way for novel diagnostic and therapeutic interventions.
At Nucleome Informatics, we are dedicated to delivering accurate and insightful lncRNA sequencing results that fuel groundbreaking discoveries in genomics research. With our advanced technologies and experienced team, we aim to be your partner in advancing the understanding of lncRNA functions across diverse research fields.
Contact us today to discuss your lncRNA sequencing project and leverage our expertise in driving transformative discoveries.


Single Cell RNA Sequencing Services at Nucleome Informatics
Nucleome Informatics now offers state-of-the-art Single Cell RNA Sequencing (scRNA-Seq) services powered by Illumina Single Cell 3 RNA Prep Kit and NovaSeq 6000 sequencing system, enabling ultra-scalable, high-resolution transcriptomics for diverse biological applications.
Innovative Single-Cell RNA Workflow
The Illumina Single Cell 3 RNA Prep kit eliminates the need for complex microfluidic instruments by introducing a vortex-based PIPseq (Particle-Templated Instant Partitioning) chemistry. This novel approach captures individual cells into emulsified partitions containing barcoded oligonucleotides on hydrogel beads. Messenger RNA (mRNA) from each cell is then isolated, barcoded, and reverse transcribed to generate unique cDNA libraries for downstream Illumina sequencing.
The workflow is streamlined into seven simple steps:
-
Cell suspension preparation
-
Cell capture with PIPseq chemistry
-
Lysis and mRNA capture
-
cDNA synthesis
-
Library preparation
-
Sequencing on NovaSeq 6000
-
Data analysis using Illumina DRAGEN Single Cell Pipeline and Partek Flow
This accessible benchtop workflow enables both new and experienced researchers to perform scRNA-Seq with high reproducibility and minimal technical barriers.
Highlights of the Technology
-
Detects high numbers of transcripts per cell, capturing delicate or rare cell types.
-
Processes from hundreds to hundreds of thousands of cells, offering scalability suited to tissue-level and cell atlas projects.
-
Compatible with methanol-fixed samples, enabling flexible experiment design across time points or remote collection sites.
-
High sensitivity and low background noise reduce sequencing artifacts and maximize biological signal clarity.
-
Highest throughput single-cell sequencing in India on NovaSeq 6000
-
End-to-end workflow from cell capture to publication-ready analysis
-
Expert bioinformatics interpretation for differential expression, clustering, and pathway enrichment
-
Flexible project design for both small-scale and population-level studies
Kits available for throughput ranging from 2,000 (T2) to 100,000 (T100) cells per sample.
-
Single Cell RNA Sequencing
Sample Requirements
Input sample should be a single-cell suspension with a viable cell count depending on the kit size selected:
-
T2 kit: Requires approximately 5,000 input cells to recover 2,000 high-quality single cells.
-
T10 kit: Requires ~17,000 input cells to recover 10,000 single cells.
-
T20 kit: Requires ~40,000 input cells for 20,000 single cells.
-
T100 kit: Requires ~200,000 input cells for up to 100,000 single cells.
The assay supports fresh or methanol-fixed cells, suitable for flexible experimental workflows including time-course or remote sample collections.
Cells are suspended and processed using Illumina Single Cell 3 RNA Prep chemistry which captures mRNA without requiring specialized microfluidic equipment.
Sequencing and Data Analysis
Using NovaSeq 6000, Nucleome delivers unmatched sequencing depth and data quality.
-
Each NovaSeq SP to S4 flow cell supports billions of reads, enabling dense sampling of up to 100,000 single cells per run.
-
Output per flow cell can reach 26 billion reads, ensuring comprehensive transcript coverage across diverse cell populations.
-
Sequencing depth is generally targeted at 20,000 reads per captured cell for optimal transcriptome coverage.
-
Typical output per sample depends on kit size and desired cell number: PLEASE SEE THE TABLE.
-
Post-sequencing analysis includes cell clustering, transcript quantification, and differential gene expression profiling, generating ready-to-use feature-barcode matrices and publication-quality data visualizations.
Applications
With Illumina’s robust chemistry and Nucleome’s analytical expertise, this platform is ideal for:
-
Cancer research: Identifying tumor cell heterogeneity and immune escape mechanisms.
-
Immunology: Mapping immune cell differentiation and pathway activation.
-
Neuroscience: Profiling complex brain tissues and discovering novel neuronal subtypes.
-
Developmental biology: Charting dynamic transcriptional programs during embryogenesis.
-
Multiomics integration: Incorporating transcriptomics with proteomics or epigenomics from the same sample.

circularRNA Sequencing
Nucleome Informatics is a leading service provider of circular RNA Sequencing in India. The circRNA digitalization analysis based on HiSeq high-throughput sequencing uses the SBS-sequencing by synthesis, which can decrease the loss of nucleotides caused by the secondary structure. It is also powerful for its small requirement of sample quantity, high through-put, high accuracy with easily operated automatic platform.
Sample requirement
- Total RNA amount: ≥ 2.0 μg; RNA concentration: ≥ 50 ng/μl (For human and mouse, 200 ng could be accepted with risk.)
- RIN value ≥ 6.3 for plants and fungi; RIN value ≥ 6.8 for animals
- OD260/280 ≥ 2.0, OD260/230 ≥ 2.0, without degradation and DNA contamination
Recommended Sequencing Depth
- ≥ 20 M reads per sample of 2x150bp on HiSeq 2500
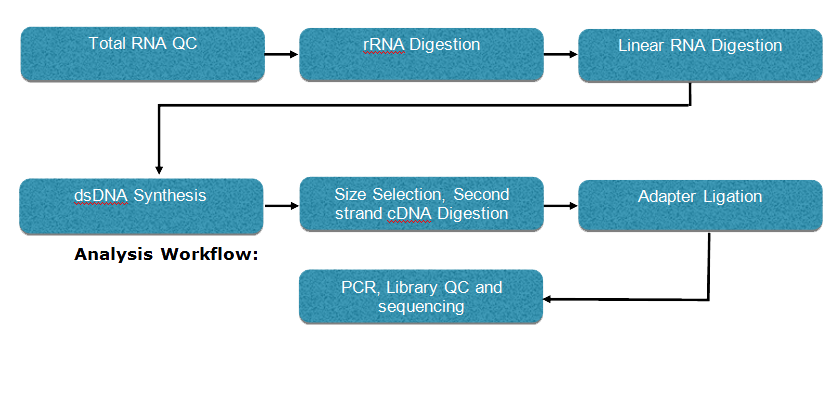
Sequencing Workflow:
The experimental procedure of circRNA sequencing is shown below:
Deliverables:
Option 1: Without Bioinformatics Analysis
- Sample QC report
- Raw Data in .fastQ format
- Raw data Stats
- Clean data QC
- Sequencing and raw data QC report
Option 2: With Bioinformatics Analysis
- Sample QC report
- Raw Data in .fastQ format
- Raw data Stats
- Clean data QC
- Sequencing and raw data QC report
- Alignment of clean reads over reference genome
- circRNA Identification, Quantification and DGE
- circRNA source gene analysis
- GO enrichment analysis
- KEGG enrichment analysis
- miRNA Binding site analysis
- Tissue specific analysis
TAT: 3 to 4 months from the date samples pass QC
DrSeq GEScore Biostimulants Testing
Biostimulants:
Biostimulants are a wide range of natural or synthetic products containing substances and microorganisms that can stimulate plant processes to improve nutrient uptake, nutrient efficiency, tolerance to abiotic stress, and crop quality.
(http:// www.biostimulants.eu/, accessed September 27, 2017)
The use of biostimulants is proposed as an advanced solution to face the demand for sustainable agriculture by ensuring optimal crop performances and better resilience to environmental changes. While the utility of some biostimulants has been proven scientifically, many growers remain sceptical of the agronomic claims.
DrSeq GEScoreTM service:
The Nucleome Informatics has developed a sequencing and data analysis approach to predict and characterise the function of natural compounds as biostimulants on plant system using DrSeq tool and BigData Decoder technology. We are the leading company offering Biostimulant testing in India based on DrSeq GEScoreTM service.
Benefits:
By using DrSeq GEScoreTM service available at Nucleome Informatics, scientists can combine plant growth assessments and transcriptomic approaches to investigate and understand the specific mode(s) of action of BIOSTIMULANTS. By using this technology, organisations manufacturing Biostimulants can decipher the scientific background how the product stimulates the growth of the plant and reduces the plant’s biotic and abiotic stress. The company can also study the change in gene expression of the plants due to a variable dose of Biostimulants or change in treatment, duration, application, location, weather, germplasm etc. This study will help the R&D teams not only in novel product development but improving the older products too.
Deliverables:
We will provide the accession number of the data submitted in NCBI and DrSeq GEScoreTM based on exact scientific parameters of the Biostimulant testing with project report which can be used to publish these results as a research paper too.
Contact Now:
For proposal, please contact our business development team at info@nucleomeinfo.com with your specific questions or call our technical team at 040 42604142 for any technical queries.



